The 3mm, 5mm, 7mm, and 9mm images are derived from the original 1mm images by resampling along the Z (slice) axis only. In order to emulate a perfectly continuous phantom (instead of one discretized into blocks) the image intensity that has been recorded experimentally for each voxel is assumed to be the image intensity at the centre of each voxel. Linear interpolation is then used to join these central intensities. The resampling is achieved as an integration of this "continuous" image (this is achieved in the simulation by effectively allowing the mixture of the tissue classes to vary in a linear fashion between voxels). Finally, when the data is ordered in space the centre of the first voxel (no matter what size the voxels are) is defined to be at z = 0.
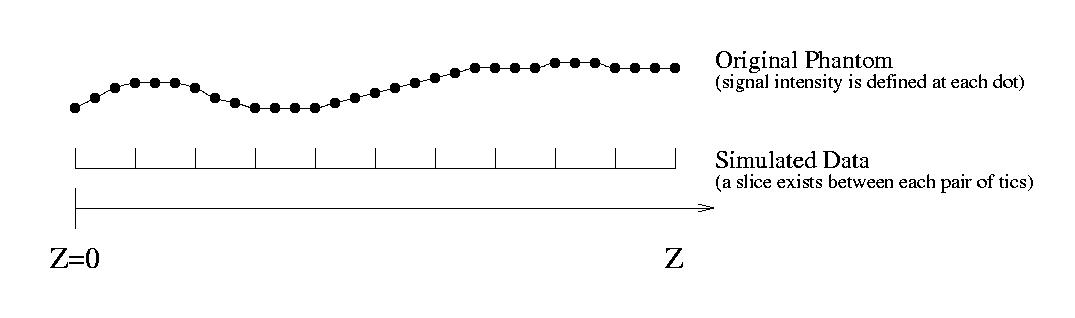
The following graph illustrates the interpolation process:
